Ergatis
Ergatis is a web-based utility that is used to create, run, and monitor reusable computational analysis pipelines. It contains pre-built components for common bioinformatics analysis tasks. These components can be arranged graphically to form highly-configurable pipelines. Each analysis component supports multiple output formats, including the Bioinformatic Sequence Markup Language (BSML). The current implementation includes support for data loading into project databases following the CHADO schema, a highly normalized, community-supported schema for storage of biological annotation data.
Ergatis uses the Workflow engine to process its work on a compute grid. Workflow provides an XML language and processing engine for specifying the steps of a computational pipeline. It provides detailed execution status and logging for process auditing, facilitates error recovery from point of failure, and is highly scalable with support for distributed computing environments. The XML format employed enables commands to be run serially, in parallel, and in any combination or nesting level.
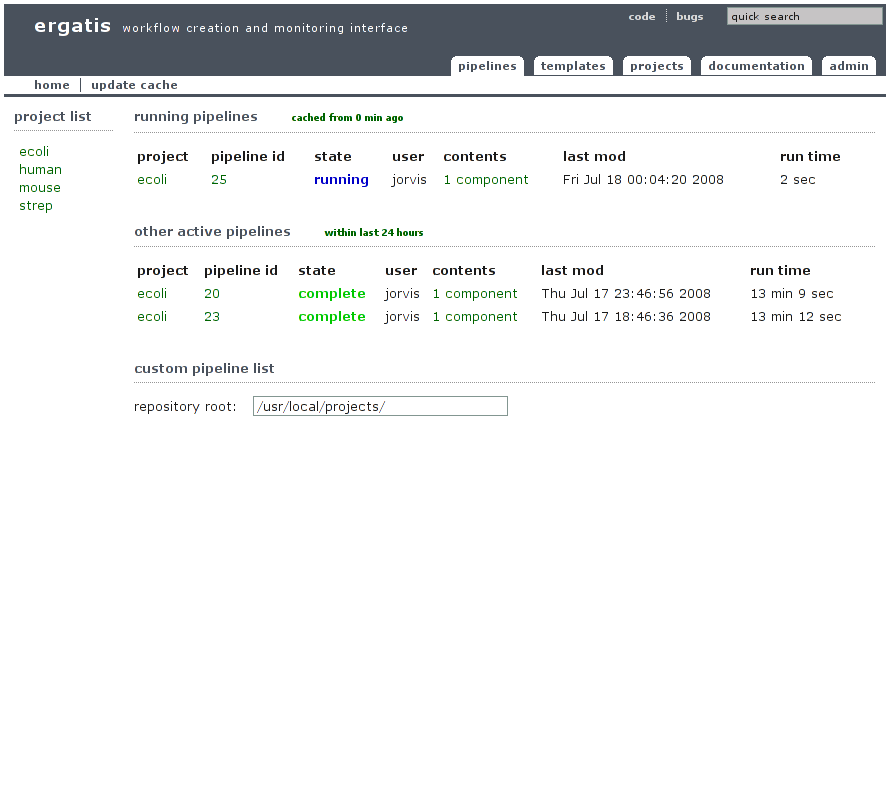
This is the main/index page for Ergatis. From it you can choose a project page or view recent pipelines run across all project.